Recently, I shared our plans for a new file format, Lance v2. As I'm creating a file reader for this new format I plan to create a series of posts talking about the design, challenges and limitations in many existing file readers, and how we plan to overcome these. Much of this discussion is not specific to Lance and I hope this series will be useful to anyone that wants to know more about storage and I/O.
In the first post of this series I want to talk about reading files in parallel. Traditionally, this has been done with row groups or stripes. In Lance v2 we got rid of row groups so we'll need to solve this in another way. Fortunately, it turns out that another way is not only possible, but actually delivers better performance, and is probably even a better way to read files in existing formats (e.g. parquet / orc).
Our First Parallel File Reader
At a high level, file readers for columnar formats often look very similar, in fact, a very similar approach was taken with Lance v1:
parallel for row_group in file:
arrays = []
parallel for column in row_group:
array = ArrayBuilder()
for page in column:
page = read_page()
array.extend(page.decode())
arrays.append(array)
batch = RecordBatch(arrays)
yield batch
This is just pseudo code of course. The real algorithm is often quite different. The decode might not happen in parallel, the page readers might have read ahead, batches might be yielded as as soon as one page is loaded for each column (instead of buffering an entire row group). These are all important details (and we may talk about them later) but not relevant to the discussion at hand. The main point is that parallelism primarily comes from accessing the row groups in parallel. This is a very intuitive way to handle parallel reading of a file. However, we got rid of row groups in Lance, and so this approach won't work for us.
Decode Based Parallelism
The solution is quite simple, I decouple my CPU batch size from my I/O read size. A single page of data actually contains quite a few values. For example, a single 1MiB page of integer data will contain at least 250,000 values. We typically want our CPU batch size to be tied to our CPU cache size. This is often 1MiB or less. If I am reading in 10 columns of data, and I have 1MiB pages then, even without any compression, I have 10 batches of data in every page.
If I'm reading a single column of data then I actually end up with the same thing I get with row groups which is one record batch / CPU task for every single data page. This ability to generate batches on a per-page basis instead of a per-group basis is useful (keeps RAM usage down) and is something that sophisticated parquet readers tend to do as well (though I'm not aware of any that apply parallelism here).
Which Approach is Better?
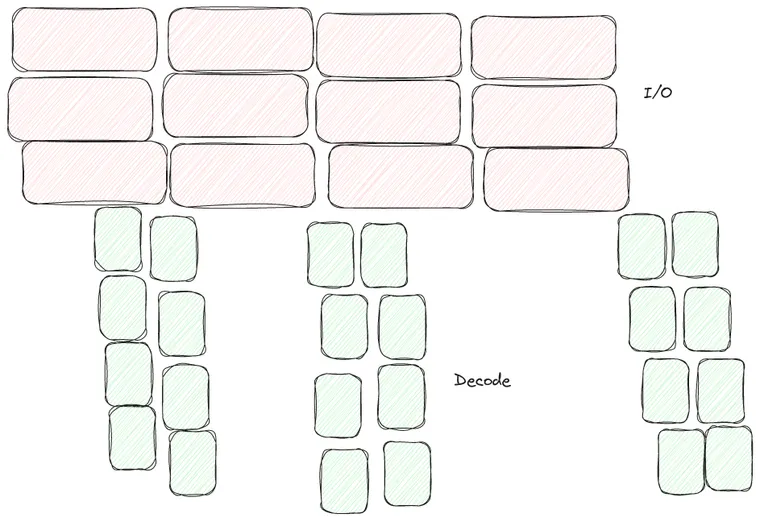
I believe the approach we are taking in Lance is not only compatible, but can actually result in better performance. The main difference is that the classic algorithm leads to the following I/O order:
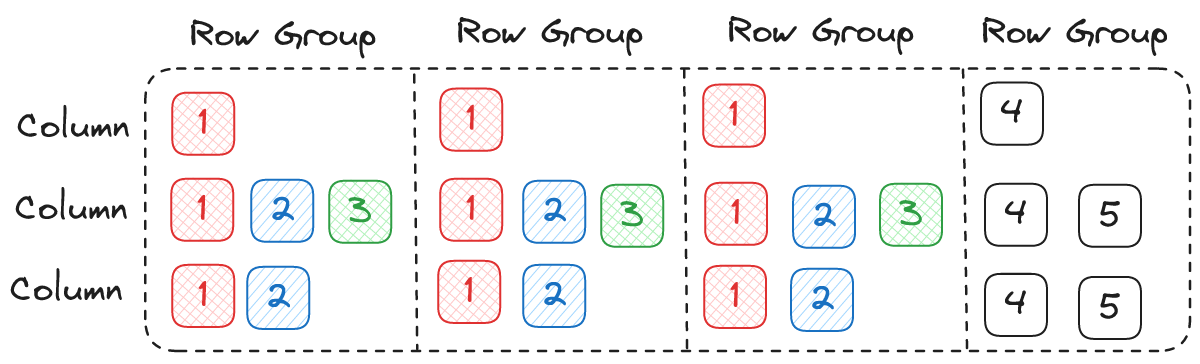
The problem with the classic reading order described above is that if you want parallelism X and you are reading Y columns then you need to perform X * Y reads in parallel. Does this matter? I/O is slow and asynchronous, we can launch as many I/O tasks in parallel as we want right?
The Myth of Infinite Parallelism
Modern disks have a lot of parallelism but they do not have infinite parallelism. The OS does an amazing job of load balancing many concurrent requests and so we typically don't care or notice this fact. In fact, over scheduling the disk slightly can be beneficial since you avoid stuttering. However, over scheduling the disk too much turns out to be a problem. To see why, let's look at some actual trace data.

Here we read the same file, with the same chunk size, but using different amounts of parallelism. I tested with 128 8-MiB chunks, across both 64 and 16 threads with a Samsung SSD 970 EVO Plus 2TB. Benchmarks report that this disk caps out at around a queue depth of 16 and so it should come as no surprise that both operations took the same amount of time since both allow for at least 16 parallel reads.
Why is this a problem?
In both cases we got the exact same bandwidth, right about 3GiBps, so this shouldn't be a problem right? However, the average request latency in the first example (64 threads) is much higher than the average latency in the second example (16 threads). Look at our parquet example again and let's consider when we can start outputting record batches. In order to return a record batch we need at least one page from each column. This means, on average, we won't be able to start emitting data until about 9 IOPS have completed.
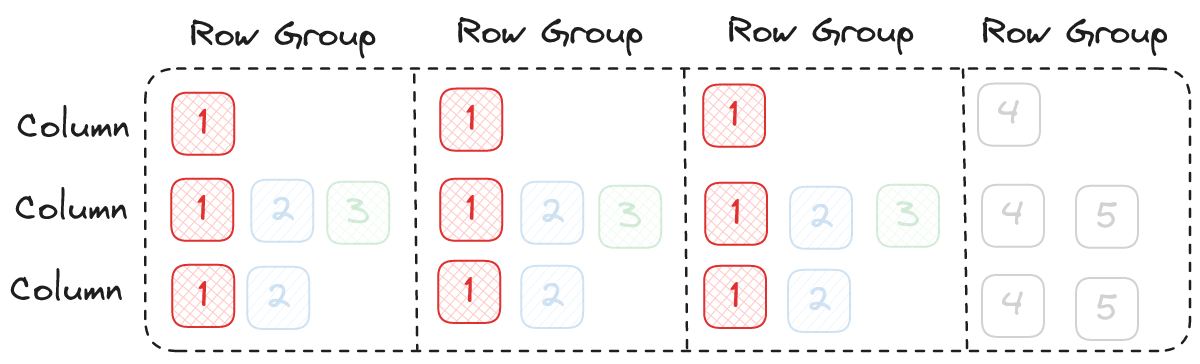
What does that mean for our file reader? It means there is a longer delay between reading and actually decoding data. This kind of delay is always necessary to some degree, but we should try to minimize it as much as possible.
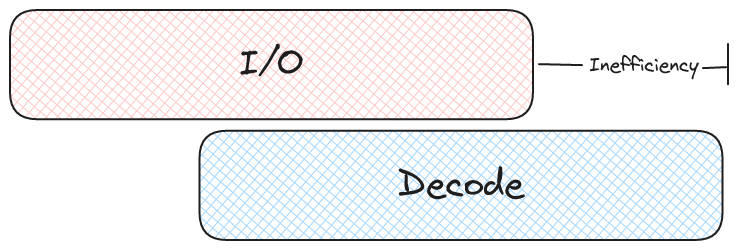
What about Cloud Storage?
Cloud storage often likes to boast about infinite I/O that can scale up to whatever bandwidth you need. However, the scaling your cloud provider is talking about it typically the number of servers and not the number of threads. This is especially true when we're talking about concurrent accesses to the same file. For example, most EC2 instances come with 1.25GiBps of network bandwidth. Here is a quote from S3's performance guide (emphasis mine):
Make one concurrent request for each 85–90 MB/s of desired network throughput. To saturate a 10 Gb/s network interface card (NIC), you might use about 15 concurrent requests over separate connections.
Even if your reads are small (this is often the case with point lookups) we have found, through practice, that the same patterns tend to hold. Too many concurrent requests and the average latency grows. We can try buying our way out of this problem by using a network optimized instance. For example, a c7gn.8xlarge will give us 10x more network bandwidth. However, we now have 32 cores, so we probably need to read even more row groups in parallel, which means that we need even more concurrent I/O. It turns out that coupling between I/O and compute parallelism is coming back to bite us.
Ideal Read Order
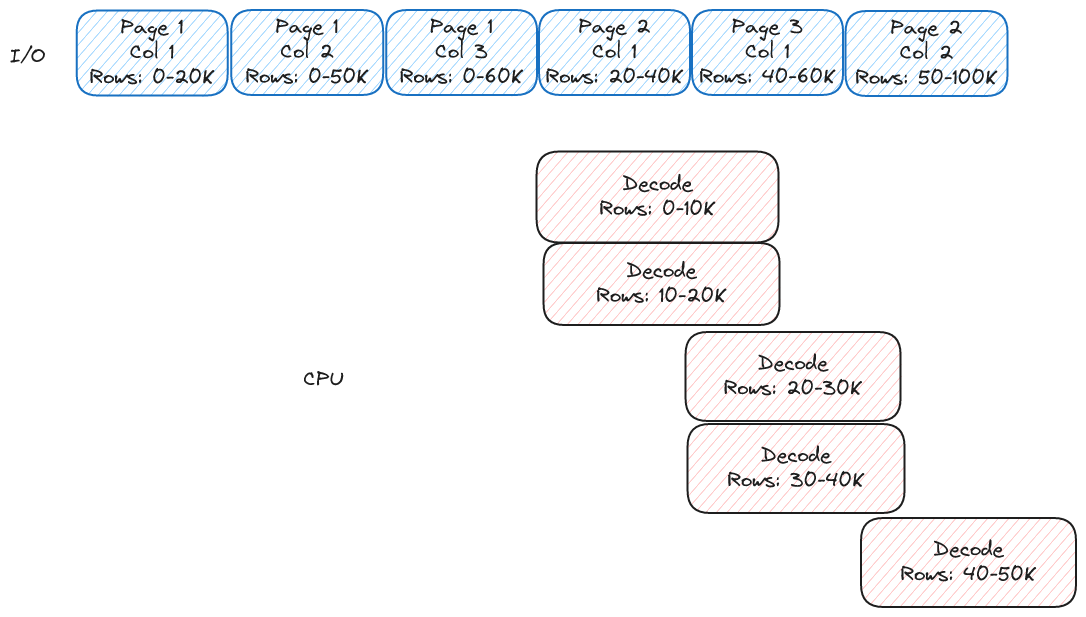
Now, let's begin talking about our solution. First, let's look at the ideal order to read our file:
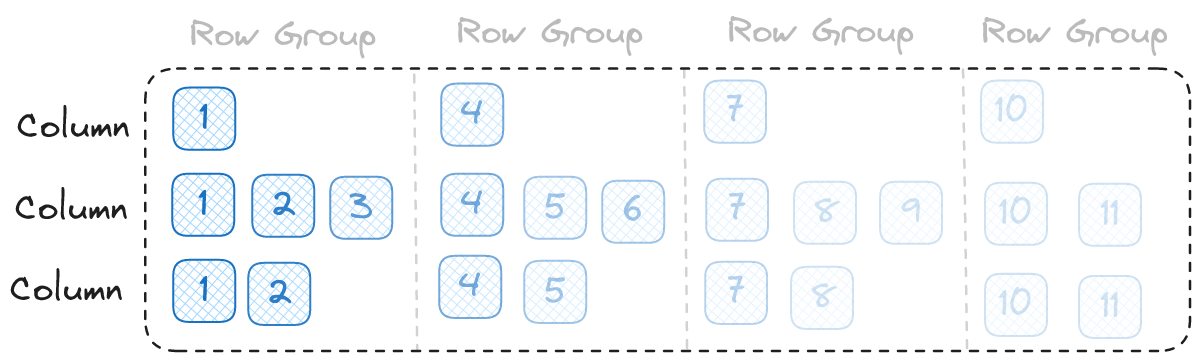
It turns out to be very simple. We read pages starting with the first row and moving up to the last row. If our ideal I/O parallelism was 6 in the above example we would read one row group at a time. If it was 12 we would read 2 row groups at a time. If it was 3 we can even read 3 pages at a time and everything will work.
Observations in Practice
So far, this discussion has been very theoretical (though it is based on issues we saw with Lance v1). Does it actually work? I think we've done enough internal benchmarking that I am convinced it does. When doing full scans of the NYC taxi dataset or the common crawl dataset the new Lance v2 format is considerably faster than Lance v1. When running locally against my NVME at 3GBps we are still I/O bound and the full read + decode bandwidth is usually within 95% of my disk's bandwidth.
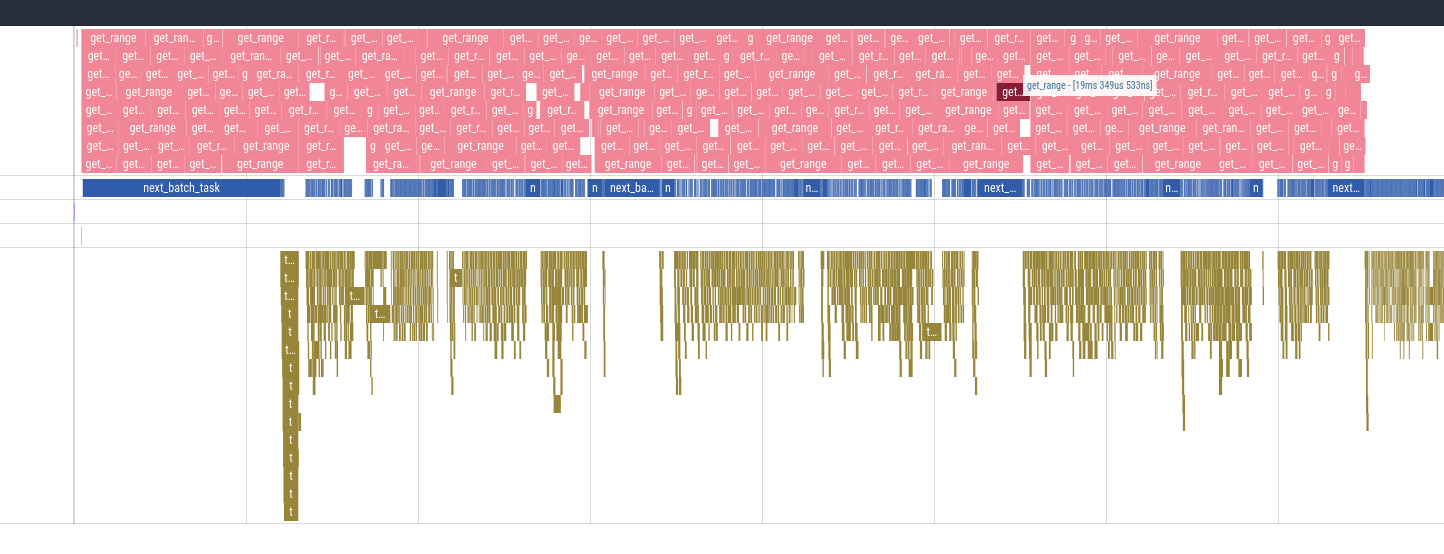
This is exciting for me as saturating my local NVME disk's bandwidth has been one of my goals with the reader.
Keep Reading
The next post in our series on file readers is now available. We talk about scheduling vs. decoding and why we separate the two processes in Lance.