We've been talking for a while about a new iteration of our file format. We're pleased to announce that the new v2 format is now in available as an opt-in feature for Lance (in release 0.12.1) and LanceDB (release 0.8.2) for users that want to try it out ahead of time and give us some early feedback.
How can you help?
We're eager for current Lance and LanceDB users to run experiments and benchmarks with the new format. We especially want to know:
- Do you run into any bugs using the new file format in your workflow?
- Are there any situations where the performance of the new file format is slower than the old format?
If you run into any issues please make sure to file an issue or let us know on Discord.
How to enable Lance v2
Turning on support for the new format is easy, but limited to creating new tables. Migration of old data will arrive as we are more confident in the stability of the format. To enable the new format simply set the use_legacy_format option to False when creating the table.
# By default, tables are still created in v1 mode
v1_tbl = await db.create_table("v1", data)
# The use_legacy_format flag will let you opt-in to v2 files
v2_tbl = await db.create_table("v2", data, use_legacy_format=False)What's available in the beta?
The beta supports all existing dataset operations today (except dictionary encoding). In addition, most of the new functionality is already available. This means you can already start taking advantage of the following features.
❓ Support for nulls (not placeholders) in all data types (except struct)
In Lance v1 we automatically converted nulls to a placeholder value (0 for numeric types and empty lists/strings for string/list types). This is a problem since it is impossible to distinguish between a null and a placeholder after the data is written. We now support nulls for all data types except struct (which we will be working on shortly).
data = pa.table(
{
"ints": [1, None],
"floats": [1.1, None],
"strings": ["", None],
"lists": [[1, 2, 3], None],
"vectors": pa.array([[1.0, 2.0], None], pa.list_(pa.float64(), 2)),
}
)
def print_table(tbl):
for name in tbl.column_names:
print(name, tbl.column(name).to_pylist())
v1_tbl = await db.create_table("v1", data)
print("With legacy format")
print_table(await v1_tbl.query().to_arrow())
# With legacy format
# ints [1, 0]
# floats [1.1, 0.0]
# strings [None, None]
# lists [[1, 2, 3], []]
# vectors [[1.0, 2.0], [0.0, 0.0]]
v2_tbl = await db.create_table("v2", data, use_legacy_format=False)
print("With new v2 format")
print_table(await v2_tbl.query().to_arrow())
# With new v2 format
# ints [1, None]
# floats [1.1, None]
# strings ['', None]
# lists [[1, 2, 3], None]
# vectors [[1.0, 2.0], None]💾 Easily create files without using excessive RAM
In Lance v1 both the RAM used by the file readers/writers and the performance of the resulting file were controlled by the row group size. In Lance v2 there is no row group size. It's now much easier to create large multi-modal datasets without using an excessive amount of memory.
db = await lancedb.connect_async("/tmp/my_db")
image_paths = glob.glob("/home/pace/dev/data/Dogs/**/*.jpg")
def load_images():
for idx, image_path in enumerate(image_paths):
img = Image.open(image_path)
img_bytes = io.BytesIO()
img.save(img_bytes, format="PNG")
yield pa.table({"img": [img_bytes.getvalue()], "ids": [idx]})
# With the v1 format this command could potentially run out of memory
# with large images or videos as it had to buffer an entire row group.
#
# This is especially true when you have many parallel writers which is
# common when your ingestion path needs to encode or decode images.
#
# With the v2 format there is no concern about RAM.
tbl = await db.create_table(
"images",
load_images(batch_size),
schema=pa.schema([pa.field("img", pa.binary()), pa.field("ids", pa.int64())]),
use_legacy_format=False,
)
# Since we don't need to clump data into small row groups any more the
# performance of this query is 6x faster than the v1 counterpart.
await tbl.query().select(["ids"]).to_arrow()🚀 Faster scans
LanceDB has always supported lightning fast vector searches. However, with the Lance v1 format, full scans of the dataset could sometimes be slow, especially when smaller row groups were needed. In Lance v2 a full dataset scan is much more efficient, giving stellar performance for scalar columns and vector columns alike.
db = await lancedb.connect_async("/tmp/my_db")
data = pq.read_table("/home/pace/dev/data/lineitem_10.parquet")
tbl = await db.create_table("lineitem", data, use_legacy_format=False)
# Querying TPC-H data is over 3x faster with Lance v2
async for batch in await tbl.query().to_batches(max_batch_length=8192):
passWhats still in progress?
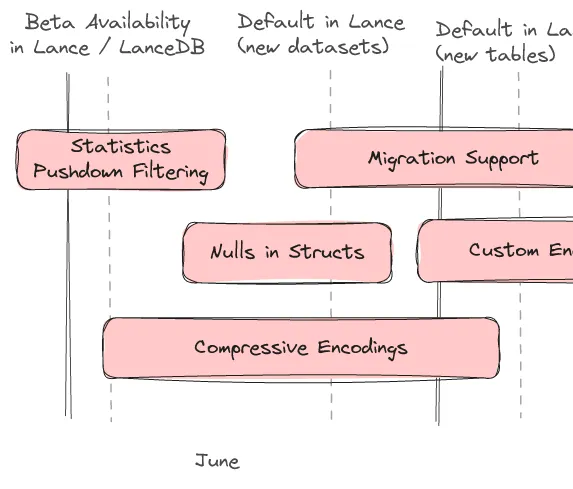
We have a rough roadmap describing the major features we will be working on over the summer.
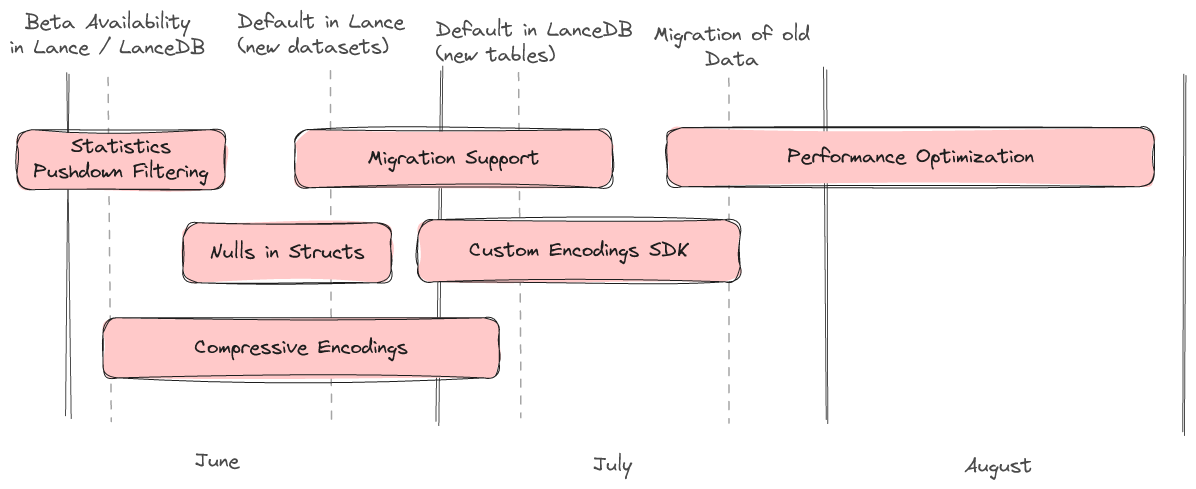
Statistics / Pushdown Filtering
Pushdown filtering is nearly complete. Lance v2 will have the same great pushdown filtering experience as Lance v1, enabling both statistics-based zone skipping as well as late materialization. Lance v2 also allows new statistics to be added as custom encodings paving the way for interesting experiments and specialized use cases.
Nulls in Structs
Null support for structs is planned and will be arriving soon. This will support "null struct", "struct of nulls", and everything in between.
Compressive Encodings
One of the most asked for features in LanceDB is the ability to use compression on scalar columns. We are in the middle of developing a number of special encodings which provide compression without slowing down point lookups or introducing excess read amplificationThis will not only lead to less disk utilization but also should lead to significantly faster queries.
Migration Support
Once the v2 format is stable we will offer a command to migrate an existing dataset to the new format.
Custom Encoding SDK
Users with specialized use cases will be able to implement their own encodings. Some examples of experiments we would like to do with this capability are "packed structs" (virtually combine several columns into one to speed up lookups that always load the same set of columns), "variants" (columns that can have more than one data type, essential for storing semi-structured or unstructured data), and multi-modal specific encodings (e.g. pushdown filtering for audio/images)
Performance Optimization
Finally, we will continue to optimize v2 at high scale in the coming weeks to further improve performance for a wide variety of use cases. If you're interested in kicking the tires on your data, we'd love to get your feedback.